cytoflowgui.op_plugins.color_translation¶
Translate measurements from one color’s scale to another, using a two-color or three-color control.
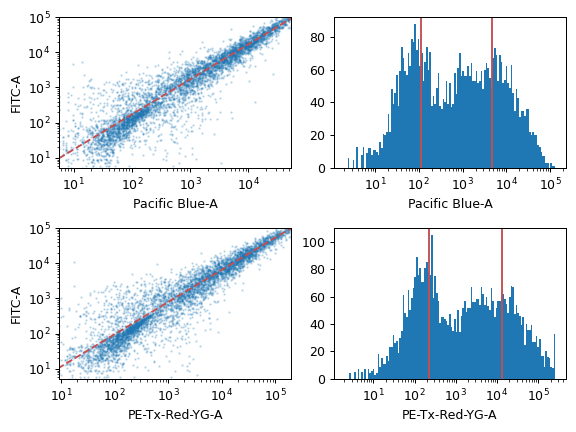
To use, set up the Controls list with the channels to convert and the FCS files to compute the mapping. Click Estimate and make sure to check that the diagnostic plots look good.
- Add Control, Remove Control
Add and remove controls to compute the channel mappings.
- Use mixture model?
If
True, try to model the from channel as a mixture of expressing cells and non-expressing cells (as you would get with a transient transfection), then weight the regression by the probability that the the cell is from the top (transfected) distribution. Make sure you check the diagnostic plots to see that this worked!
Note
You cannot have any operations before this one which estimate model parameters based on experimental conditions. (Eg, you can’t use a Density Gate to choose morphological parameters and set by to an experimental condition.) If you need this functionality, you can access it using the Python module interface.
- class cytoflowgui.op_plugins.color_translation.ControlHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.Controller
- class cytoflowgui.op_plugins.color_translation.ColorTranslationHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
cytoflowgui.op_plugins.op_plugin_base.OpHandler- add_control¶
alias of
traits.trait_types.Event
- remove_control¶
alias of
traits.trait_types.Event
- channels = <traits.traits.ForwardProperty object>¶
- class cytoflowgui.op_plugins.color_translation.ColorTranslationViewHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
cytoflowgui.view_plugins.view_plugin_base.ViewHandler
- class cytoflowgui.op_plugins.color_translation.ColorTranslationPlugin[source]¶
Bases:
envisage.plugin.Plugin,cytoflowgui.op_plugins.op_plugin_base.PluginHelpMixin- operation_id = 'edu.mit.synbio.cytoflow.operations.color_translation'¶
- view_id = 'edu.mit.synbio.cytoflow.view.colortranslationdiagnostic'¶
- short_name = 'Color Translation'¶