cytoflowgui.op_plugins.bleedthrough_linear¶
Apply matrix-based bleedthrough correction to a set of fluorescence channels.
This is a traditional matrix-based compensation for bleedthrough. For each pair of channels, the module estimates the proportion of the first channel that bleeds through into the second, then performs a matrix multiplication to compensate the raw data.
This works best on data that has had autofluorescence removed first; if that is the case, then the autofluorescence will be subtracted from the single-color controls too.
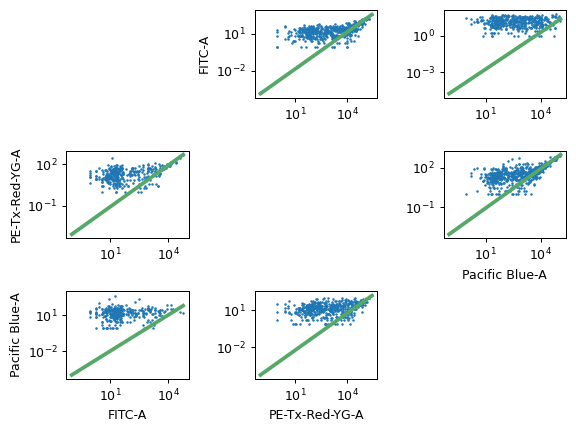
To use, specify the single-color control files and which channels they should be measured in, then click Estimate. Check the diagnostic plot to make sure the estimation looks good. There must be at least two channels corrected.
- Add Control, Remove Control
Add or remove single-color controls.
Note
You cannot have any operations before this one which estimate model parameters based on experimental conditions. (Eg, you can’t use a Density Gate to choose morphological parameters and set by to an experimental condition.) If you need this functionality, you can access it using the Python module interface.
- class cytoflowgui.op_plugins.bleedthrough_linear.ControlHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.Controller
- class cytoflowgui.op_plugins.bleedthrough_linear.BleedthroughLinearHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
cytoflowgui.op_plugins.op_plugin_base.OpHandler- add_control¶
alias of
traits.trait_types.Event
- remove_control¶
alias of
traits.trait_types.Event
- channels = <traits.traits.ForwardProperty object>¶
- class cytoflowgui.op_plugins.bleedthrough_linear.BleedthroughLinearViewHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
cytoflowgui.view_plugins.view_plugin_base.ViewHandler
- class cytoflowgui.op_plugins.bleedthrough_linear.BleedthroughLinearPlugin[source]¶
Bases:
envisage.plugin.Plugin,cytoflowgui.op_plugins.op_plugin_base.PluginHelpMixin- operation_id = 'edu.mit.synbio.cytoflow.operations.bleedthrough_linear'¶
- view_id = 'edu.mit.synbio.cytoflow.view.linearbleedthroughdiagnostic'¶
- short_name = 'Linear Compensation'¶