cytoflowgui.op_plugins.gaussian_2d¶
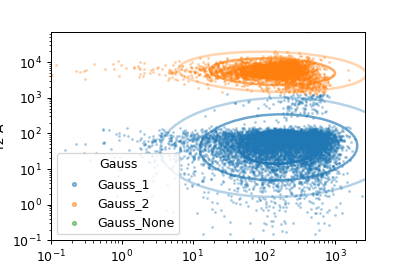
Fit a Gaussian mixture model with a specified number of components to two channels.
If Num Components is greater than 1, then this module creates a new
categorical metadata variable named Name, with possible values
{name}_1 …. name_n where n is the number of components.
An event is assigned to name_i category if it has the highest posterior
probability of having been produced by component i. If an event has a
value that is outside the range of one of the channels’ scales, then it is
assigned to {name}_None.
Additionally, if Sigma is greater than 0, this module creates new boolean
metadata variables named {name}_1 … {name}_n where n is the
number of components. The column {name}_i is True if the event is less
than Sigma standard deviations from the mean of component i. If
Num Components is 1, Sigma must be greater than 0.
Finally, the same mixture model (mean and standard deviation) may not be appropriate for every subset of the data. If this is the case, you can use By to specify metadata by which to aggregate the data before estimating and applying a mixture model.
Note
Num Components and Sigma will be the same for each subset.
- Name
The operation name; determines the name of the new metadata
- X Channel, Y Channel
The channels to apply the mixture model to.
- X Scale, Y Scale
Re-scale the data in Channel before fitting.
- Num Components
How many components to fit to the data? Must be a positive integer.
- Sigma
How many standard deviations on either side of the mean to include in the boolean variable
{name}_i? Must be>= 0.0. If Num Components is1, must be> 0.
- By
A list of metadata attributes to aggregate the data before estimating the model. For example, if the experiment has two pieces of metadata,
TimeandDox, setting By to["Time", "Dox"]will fit the model separately to each subset of the data with a unique combination ofTimeandDox.
- class cytoflowgui.op_plugins.gaussian_2d.GaussianMixture2DHandler(*args: Any, **kwargs: Any)[source]¶
- class cytoflowgui.op_plugins.gaussian_2d.GaussianMixture2DViewHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
cytoflowgui.view_plugins.view_plugin_base.ViewHandler
- class cytoflowgui.op_plugins.gaussian_2d.GaussianMixture2DPlugin[source]¶
Bases:
envisage.plugin.Plugin,cytoflowgui.op_plugins.op_plugin_base.PluginHelpMixin- operation_id = 'edu.mit.synbio.cytoflowgui.operations.gaussian_2d'¶
- view_id = 'edu.mit.synbio.cytoflow.view.gaussianmixture2dview'¶
- short_name = '2D Mixture Model'¶