cytoflowgui.op_plugins.kmeans¶
This module uses the KMeans algorithm to assign events to clusters in an unsupervized manner.
- Name
The operation name; determines the name of the new metadata
- X Channel, Y Channel
The channels to apply the mixture model to.
- X Scale, Y Scale
Re-scale the data in Channel before fitting.
- Num Clusters
How many clusters to assign the data to.
- By
A list of metadata attributes to aggregate the data before estimating the model. For example, if the experiment has two pieces of metadata,
TimeandDox, setting By to["Time", "Dox"]will fit the model separately to each subset of the data with a unique combination ofTimeandDox.
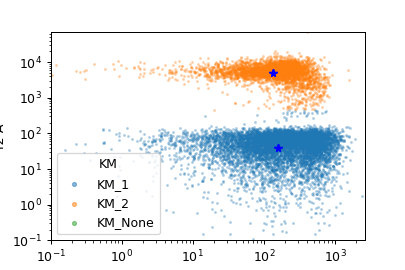
- class cytoflowgui.op_plugins.kmeans.KMeansHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.
- class cytoflowgui.op_plugins.kmeans.KMeansViewHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.
- class cytoflowgui.op_plugins.kmeans.KMeansPlugin[source]¶
Bases:
envisage.plugin.Plugin,cytoflowgui.op_plugins.op_plugin_base.PluginHelpMixin- operation_id = 'edu.mit.synbio.cytoflow.operations.kmeans'¶
- view_id = 'edu.mit.synbio.cytoflowgui.op_plugins.kmeans'¶
- short_name = 'KMeans'¶