cytoflowgui.op_plugins.density¶
Computes a gate based on a 2D density plot. The user chooses what proportion of events to keep, and the module creates a gate that selects those events in the highest-density bins of a 2D density histogram.
A single gate may not be appropriate for an entire experiment. If this is the case, you can use By to specify metadata by which to aggregate the data before computing and applying the gate.
- Name
The operation name; determines the name of the new metadata
- X Channel, Y Channel
The channels to apply the mixture model to.
- X Scale, Y Scale
Re-scale the data in Channel before fitting.
- Keep
The proportion of events to keep in the gate. Defaults to 0.9.
- By
A list of metadata attributes to aggregate the data before estimating the model. For example, if the experiment has two pieces of metadata,
TimeandDox, setting by to["Time", "Dox"]will fit the model separately to each subset of the data with a unique combination ofTimeandDox.
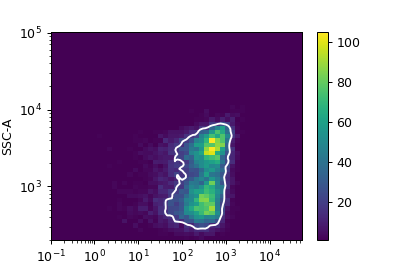
- class cytoflowgui.op_plugins.density.DensityGateHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.
- class cytoflowgui.op_plugins.density.DensityGateViewHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.
- class cytoflowgui.op_plugins.density.DensityGatePlugin[source]¶
Bases:
envisage.plugin.Plugin,cytoflowgui.op_plugins.op_plugin_base.PluginHelpMixin- operation_id = 'edu.mit.synbio.cytoflow.operations.density'¶
- view_id = 'edu.mit.synbio.cytoflow.view.densitygateview'¶
- short_name = 'Density Gate'¶