cytoflow.operations.bleedthrough_linear¶
The bleedthrough_linear module contains two classes:
BleedthroughLinearOp – compensates for spectral bleedthrough
in a Experiment using single-color controls
BleedthroughLinearDiagnostic – a diagnostic view to make sure
that BleedthroughLinearOp correctly estimated its parameters.
- class cytoflow.operations.bleedthrough_linear.BleedthroughLinearOp[source]¶
Bases:
traits.has_traits.HasStrictTraitsApply matrix-based bleedthrough correction to a set of fluorescence channels.
This is a traditional matrix-based compensation for bleedthrough. For each pair of channels, the user specifies the proportion of the first channel that bleeds through into the second; then, the module performs a matrix multiplication to compensate the raw data.
The module can also estimate the bleedthrough matrix using one single-color control per channel.
This works best on data that has had autofluorescence removed first; if that is the case, then the autofluorescence will be subtracted from the single-color controls too.
To use, set up the
controlsdict with the single color controls; callestimateto parameterize the operation; check that the bleedthrough plots look good by callingBleedthroughLinearDiagnostic.ploton theBleedthroughLinearDiagnosticinstance returned bydefault_view; and thenapplyon anExperiment.- controls¶
The channel names to correct, and corresponding single-color control FCS files to estimate the correction splines with. Must be set to use
estimate.- Type
Dict(Str, File)
- spillover¶
The spillover “matrix” to use to correct the data. The keys are pairs of channels, and the values are proportions of spectral overlap. If
("channel1", "channel2")is present as a key,("channel2", "channel1")must also be present. The module does not assume that the matrix is symmetric.- Type
Dict(Tuple(Str, Str), Float)
- control_conditions¶
Occasionally, you’ll need to specify the experimental conditions that the bleedthrough tubes were collected under (to apply the operations in the history.) Specify them here. The key is the channel name; they value is a dictionary of the conditions (same as you would specify for a
cytoflow.operations.import_op.Tube)- Type
Dict(Str, Dict(Str, Any))
Examples
Create a small experiment:
>>> import cytoflow as flow >>> import_op = flow.ImportOp() >>> import_op.tubes = [flow.Tube(file = "tasbe/rby.fcs")] >>> ex = import_op.apply()
Correct for autofluorescence
>>> af_op = flow.AutofluorescenceOp() >>> af_op.channels = ["Pacific Blue-A", "FITC-A", "PE-Tx-Red-YG-A"] >>> af_op.blank_file = "tasbe/blank.fcs"
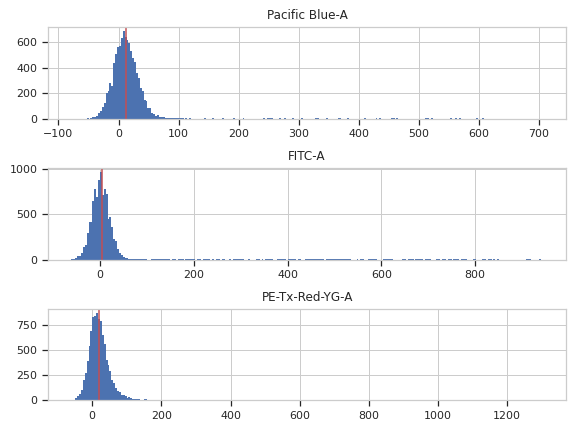
>>> af_op.estimate(ex) >>> af_op.default_view().plot(ex)
>>> ex2 = af_op.apply(ex)
Create and parameterize the operation
>>> bl_op = flow.BleedthroughLinearOp() >>> bl_op.controls = {'Pacific Blue-A' : 'tasbe/ebfp.fcs', ... 'FITC-A' : 'tasbe/eyfp.fcs', ... 'PE-Tx-Red-YG-A' : 'tasbe/mkate.fcs'}
Estimate the model parameters
>>> bl_op.estimate(ex2)
Plot the diagnostic plot
Note
The diagnostic plots look really bad in the online documentation. They’re better in a real-world example, I promise!
>>> bl_op.default_view().plot(ex2)
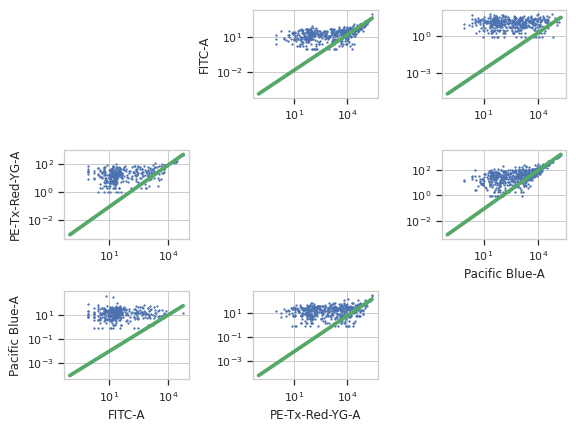
Apply the operation to the experiment
>>> ex2 = bl_op.apply(ex2)
- estimate(experiment, subset=None)[source]¶
Estimate the bleedthrough from simgle-channel controls in
controls
- apply(experiment)[source]¶
Applies the bleedthrough correction to an experiment.
- Parameters
experiment (
Experiment) – The experiment to which this operation is applied- Returns
A new
Experimentwith the bleedthrough subtracted out. The corrected channels have the following metadata added:linear_bleedthrough : Dict(Str : Float) The values for spillover from other channels into this channel.
bleedthrough_channels : List(Str) The channels that were used to correct this one.
bleedthrough_fn : Callable (Tuple(Float) –> Float) The function that will correct one event in this channel. Pass it the values specified in
controlsand it will return the corrected value for this channel.
- Return type
- default_view(**kwargs)[source]¶
Returns a diagnostic plot to make sure spillover estimation is working.
- Returns
An
IView, callBleedthroughLinearDiagnostic.plotto see the diagnostic plots- Return type
- class cytoflow.operations.bleedthrough_linear.BleedthroughLinearDiagnostic[source]¶
Bases:
traits.has_traits.HasStrictTraitsPlots a scatterplot of each channel vs every other channel and the bleedthrough line
- op¶
The operation whose parameters we’re viewing. If you made the instance with
BleedthroughLinearOp.default_view, this is set for you already.- Type
Instance(BleedthroughPiecewiseOp)