cytoflowgui.op_plugins.autofluorescence¶
Apply autofluorescence correction to a set of fluorescence channels.
This module estimates the arithmetic median fluorescence from cells that are not fluorescent, then subtracts the median from the experimental data.
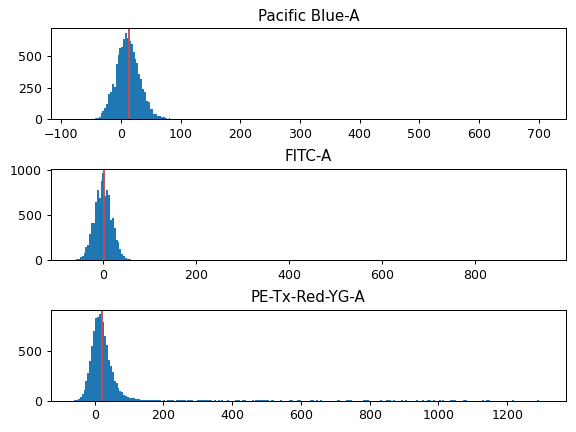
Check the diagnostic plot to make sure that the sample is actually non-fluorescent, and that the module found the population median.
- Channels
The channels to correct
- Blank file
The FCS file containing measurements of blank cells.
Note
You cannot have any operations before this one which estimate model parameters based on experimental conditions. (Eg, you can’t use a Density Gate to choose morphological parameters and set by to an experimental condition.) If you need this functionality, you can access it using the Python module interface.
- class cytoflowgui.op_plugins.autofluorescence.AutofluorescenceHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.
- class cytoflowgui.op_plugins.autofluorescence.AutofluorescenceViewHandler(*args: Any, **kwargs: Any)[source]¶
Bases:
traitsui.api.
- class cytoflowgui.op_plugins.autofluorescence.AutofluorescencePlugin[source]¶
Bases:
envisage.plugin.Plugin,cytoflowgui.op_plugins.op_plugin_base.PluginHelpMixin- operation_id = 'edu.mit.synbio.cytoflow.operations.autofluorescence'¶
- view_id = 'edu.mit.synbio.cytoflow.view.autofluorescencediagnosticview'¶
- short_name = 'Autofluorescence correction'¶