cytoflow.operations.range¶
-
class
cytoflow.operations.range.RangeOp[source]¶ Bases:
traits.has_traits.HasStrictTraitsApply a range gate to a cytometry experiment.
-
name¶ The operation name. Used to name the new metadata field in the experiment that’s created by
apply()Type: Str
-
channel¶ The name of the channel to apply the range gate.
Type: Str
-
low¶ The lowest value to include in this gate.
Type: Float
-
high¶ The highest value to include in this gate.
Type: Float
Examples
Make a little data set.
>>> import cytoflow as flow >>> import_op = flow.ImportOp() >>> import_op.tubes = [flow.Tube(file = "Plate01/RFP_Well_A3.fcs", ... conditions = {'Dox' : 10.0}), ... flow.Tube(file = "Plate01/CFP_Well_A4.fcs", ... conditions = {'Dox' : 1.0})] >>> import_op.conditions = {'Dox' : 'float'} >>> ex = import_op.apply()
Create and parameterize the operation.
>>> range_op = flow.RangeOp(name = 'Range', ... channel = 'Y2-A', ... low = 2000, ... high = 10000)
Plot a diagnostic view
>>> rv = range_op.default_view(scale = 'log') >>> rv.plot(ex)
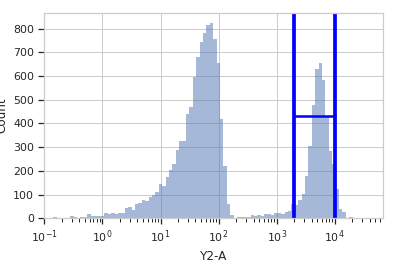
Note
If you want to use the interactive default view in a Jupyter notebook, make sure you say
%matplotlib notebookin the first cell (instead of%matplotlib inlineor similar). Then calldefault_view()withinteractive = True:rv = range_op.default_view(scale = 'log', interactive = True) rv.plot(ex)
Apply the gate, and show the result
>>> ex2 = range_op.apply(ex) >>> ex2.data.groupby('Range').size() Range False 16042 True 3958 dtype: int64
-
apply(experiment)[source]¶ Applies the range gate to an experiment.
Parameters: experiment (Experiment) – the old_experiment to which this op is applied Returns: a new experiment, the same as old Experimentbut with a new column of typeboolwith the same as the operation name. The bool isTrueif the event’s measurement inchannelis greater thanlowand less thanhigh; it isFalseotherwise.Return type: Experiment
-
-
class
cytoflow.operations.range.RangeSelection[source]¶ Bases:
cytoflow.operations.base_op_views.Op1DView,cytoflow.views.histogram.HistogramViewPlots, and lets the user interact with, a selection on the X axis.
-
interactive¶ is this view interactive? Ie, can the user set min and max with a mouse drag?
Type: Bool
-
channel¶ The channel this view is viewing. If you created the view using
default_view(), this is already set.Type: String
-
scale¶ The way to scale the x axes. If you created the view using
default_view(), this may be already set.Type: {‘linear’, ‘log’, ‘logicle’}
-
op¶ The
IOperationthat this view is associated with. If you created the view usingdefault_view(), this is already set.Type: Instance(IOperation)
-
xfacet, yfacet Set to one of the
conditionsin theExperiment, and a new row or column of subplots will be added for every unique value of that condition.Type: String
-
huefacet¶ Set to one of the
conditionsin the in theExperiment, and a new color will be added to the plot for every unique value of that condition.Type: String
Notes
We inherit
xfacetandyfacetfrom cytoflow.views.HistogramView, but they must both be unset!Examples
In an IPython notebook with %matplotlib notebook
>>> r = RangeOp(name = "RangeGate", ... channel = 'Y2-A') >>> rv = r.default_view() >>> rv.interactive = True >>> rv.plot(ex2) >>> ### draw a range on the plot ### >>> print r.low, r.high
-
plot(experiment, **kwargs)[source]¶ Plot the underlying histogram and then plot the selection on top of it.
Parameters: - experiment (Experiment) – The
Experimentto plot using this view. - title (str) – Set the plot title
- xlabel, ylabel (str) – Set the X and Y axis labels
- huelabel (str) – Set the label for the hue facet (in the legend)
- legend (bool) – Plot a legend for the color or hue facet? Defaults to True.
- sharex, sharey (bool) – If there are multiple subplots, should they share axes? Defaults to True.
- row_order, col_order, hue_order (list) – Override the row/column/hue facet value order with the given list. If a value is not given in the ordering, it is not plotted. Defaults to a “natural ordering” of all the values.
- height (float) – The height of each row in inches. Default = 3.0
- aspect (float) – The aspect ratio of each subplot. Default = 1.5
- col_wrap (int) – If xfacet is set and yfacet is not set, you can “wrap” the subplots around so that they form a multi-row grid by setting col_wrap to the number of columns you want.
- sns_style ({“darkgrid”, “whitegrid”, “dark”, “white”, “ticks”}) – Which seaborn style to apply to the plot? Default is whitegrid.
- sns_context ({“paper”, “notebook”, “talk”, “poster”}) – Which seaborn context to use? Controls the scaling of plot elements such as tick labels and the legend. Default is talk.
- despine (Bool) – Remove the top and right axes from the plot? Default is True.
- min_quantile (float (>0.0 and <1.0, default = 0.001)) – Clip data that is less than this quantile.
- max_quantile (float (>0.0 and <1.0, default = 1.00)) – Clip data that is greater than this quantile.
- lim ((float, float)) – Set the range of the plot’s data axis.
- orientation ({‘vertical’, ‘horizontal’})
- num_bins (int) – The number of bins to plot in the histogram. Clipped to [100, 1000]
- histtype ({‘stepfilled’, ‘step’, ‘bar’}) – The type of histogram to draw. stepfilled is the default, which is a line plot with a color filled under the curve.
- density (bool) – If True, re-scale the histogram to form a probability density function, so the area under the histogram is 1.
- linewidth (float) – The width of the histogram line (in points)
- linestyle ([‘-‘ | ‘–’ | ‘-.’ | ‘:’ | “None”]) – The style of the line to plot
- alpha (float (default = 0.5)) – The alpha blending value, between 0 (transparent) and 1 (opaque).
- experiment (Experiment) – The
-