cytoflow.operations.pca¶
-
class
cytoflow.operations.pca.PCAOp[source]¶ Bases:
traits.has_traits.HasStrictTraitsUse principal components analysis (PCA) to decompose a multivariate data set into orthogonal components that explain a maximum amount of variance.
Call
estimate()to compute the optimal decomposition.Calling
apply()creates new “channels” named{name}_1 ... {name}_n, wherenameis thenameattribute andnisnum_components.The same decomposition may not be appropriate for different subsets of the data set. If this is the case, you can use the
byattribute to specify metadata by which to aggregate the data before estimating (and applying) a model. The PCA parameters such as the number of components and the kernel are the same across each subset, though.-
name¶ The operation name; determines the name of the new columns.
Type: Str
-
channels¶ The channels to apply the decomposition to.
Type: List(Str)
-
scale¶ Re-scale the data in the specified channels before fitting. If a channel is in
channelsbut not inscale, the current package-wide default (set withset_default_scale()) is used.Type: Dict(Str : {“linear”, “logicle”, “log”})
-
num_components¶ How many components to fit to the data? Must be a positive integer.
Type: Int (default = 2)
-
by¶ A list of metadata attributes to aggregate the data before estimating the model. For example, if the experiment has two pieces of metadata,
TimeandDox, settingbyto["Time", "Dox"]will fit the model separately to each subset of the data with a unique combination ofTimeandDox.Type: List(Str)
-
whiten¶ Scale each component to unit variance? May be useful if you will be using unsupervized clustering (such as K-means).
Type: Bool (default = False)
Examples
Make a little data set.
>>> import cytoflow as flow >>> import_op = flow.ImportOp() >>> import_op.tubes = [flow.Tube(file = "Plate01/RFP_Well_A3.fcs", ... conditions = {'Dox' : 10.0}), ... flow.Tube(file = "Plate01/CFP_Well_A4.fcs", ... conditions = {'Dox' : 1.0})] >>> import_op.conditions = {'Dox' : 'float'} >>> ex = import_op.apply()
Create and parameterize the operation.
>>> pca = flow.PCAOp(name = 'PCA', ... channels = ['V2-A', 'V2-H', 'Y2-A', 'Y2-H'], ... scale = {'V2-A' : 'log', ... 'V2-H' : 'log', ... 'Y2-A' : 'log', ... 'Y2-H' : 'log'}, ... num_components = 2, ... by = ["Dox"])
Estimate the decomposition
>>> pca.estimate(ex)
Apply the operation
>>> ex2 = pca.apply(ex)
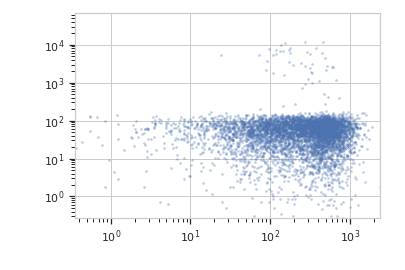
Plot a scatterplot of the PCA. Compare to a scatterplot of the underlying channels.
>>> flow.ScatterplotView(xchannel = "V2-A", ... xscale = "log", ... ychannel = "Y2-A", ... yscale = "log", ... subset = "Dox == 1.0").plot(ex2)
>>> flow.ScatterplotView(xchannel = "PCA_1", ... ychannel = "PCA_2", ... subset = "Dox == 1.0").plot(ex2)
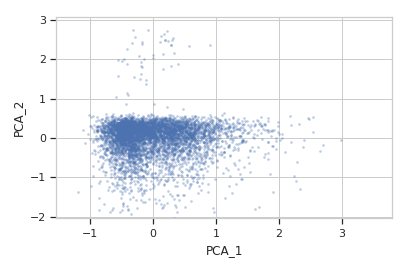
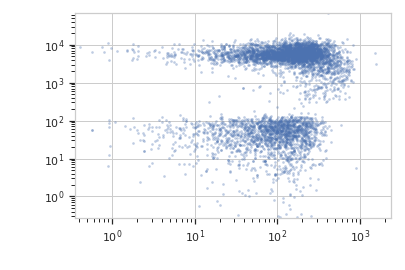
>>> flow.ScatterplotView(xchannel = "V2-A", ... xscale = "log", ... ychannel = "Y2-A", ... yscale = "log", ... subset = "Dox == 10.0").plot(ex2)
>>> flow.ScatterplotView(xchannel = "PCA_1", ... ychannel = "PCA_2", ... subset = "Dox == 10.0").plot(ex2)
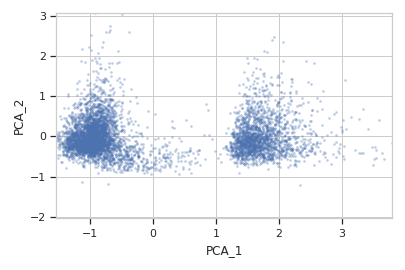
-
estimate(experiment, subset=None)[source]¶ Estimate the decomposition
Parameters: - experiment (Experiment) – The
Experimentto use to estimate the k-means clusters - subset (str (default = None)) – A Python expression that specifies a subset of the data in
experimentto use to parameterize the operation.
- experiment (Experiment) – The
-
apply(experiment)[source]¶ Apply the PCA decomposition to the data.
Returns: a new Experiment with additional channelsnamedname_1 ... name_nReturn type: Experiment
-