cytoflow.operations.bead_calibration¶
-
class
cytoflow.operations.bead_calibration.BeadCalibrationOp[source]¶ Bases:
traits.has_traits.HasStrictTraitsCalibrate arbitrary channels to molecules-of-fluorophore using fluorescent beads (eg, the Spherotech RCP-30-5A rainbow beads.)
Computes a log-linear calibration function that maps arbitrary fluorescence units to physical units (ie molecules equivalent fluorophore, or MEF).
To use, set
beads_fileto an FCS file containing events collected using the same cytometer settings as the data you’re calibrating. Specify which beads you ran by settingbeadsto match one of the values ofBeadCalibrationOp.BEADS; and setunitsto which channels you want calibrated and in which units. Then, callestimate()and check the peak-finding withdefault_view().plot(). If the peak-finding is wacky, try adjustingbead_peak_quantileandbead_brightness_threshold. When the peaks are successfully identified, callapply()to scale your experimental data set.If you can’t make the peak finding work, please submit a bug report!
This procedure works best when the beads file is very clean data. It does not do its own gating (maybe a future addition?) In the meantime, I recommend gating the acquisition on the FSC/SSC channels in order to get rid of debris, cells, and other noise.
Finally, because you can’t have a negative number of fluorescent molecules (MEFLs, etc) (as well as for math reasons), this module filters out negative values.
-
units¶ A dictionary specifying the channels you want calibrated (keys) and the units you want them calibrated in (values). The units must be keys of the
beadsattribute.Type: Dict(Str, Str)
-
beads_file¶ A file containing the FCS events from the beads.
Type: File
-
beads¶ The beads’ characteristics. Keys are calibrated units (ie, MEFL or MEAP) and values are ordered lists of known fluorophore levels. Common values for this dict are included in
BeadCalibrationOp.BEADS.Type: Dict(Str, List(Float))
-
bead_peak_quantile¶ The quantile threshold used to choose bead peaks.
Type: Int (default = 80)
-
bead_brightness_threshold¶ How bright must a bead peak be to be considered?
Type: Float (default = 100)
-
bead_brightness_cutoff¶ If a bead peak is above this, then don’t consider it. Takes care of clipping saturated detection. Defaults to 70% of the detector range.
Type: Float
-
bead_histogram_bins¶ The number of bins to use in computing the bead histogram. Tweak this if the peak find is having difficulty, or if you have a small number of events
Type: Int (default = 512)
-
force_linear¶ A linear fit in log space doesn’t always go through the origin, which means that the calibration function isn’t strictly a multiplicative scaling operation. Set
force_linearto force the such behavior. Keep an eye on the diagnostic plot, though, to see how much error you’re introducing!Type: Bool (default = False)
Notes
The peak finding is rather sophisticated.
For each channel, a 256-bin histogram is computed on the log-transformed bead data, and then the histogram is smoothed with a Savitzky-Golay filter (with a window length of 5 and a polynomial order of 1).
Next, a wavelet-based peak-finding algorithm is used: it convolves the smoothed histogram with a series of wavelets and looks for relative maxima at various length-scales. The parameters of the smoothing algorithm were arrived at empircally, using beads collected at a wide range of PMT voltages.
Finally, the peaks are filtered by height (the histogram bin has a quantile greater than bead_peak_quantile) and intensity (brighter than
bead_brightness_threshold).How to convert from a series of peaks to mean equivalent fluorochrome? If there’s one peak, we assume that it’s the brightest peak. If there are two peaks, we assume they’re the brightest two. If there are
n >=3peaks, we check all the contiguous n-subsets of the bead intensities and find the one whose linear regression (in log space!) has the smallest norm (square-root sum-of-squared-residuals.)There’s a slight subtlety in the fact that we’re performing the linear regression in log-space: if the relationship in log10-space is
Y=aX + b, then the same relationship in linear space isx = 10**X,y = 10**y, andy = (10**b) * (x ** a).Examples
Create a small experiment:
>>> import cytoflow as flow >>> import_op = flow.ImportOp() >>> import_op.tubes = [flow.Tube(file = "tasbe/rby.fcs")] >>> ex = import_op.apply()
Create and parameterize the operation
>>> bead_op = flow.BeadCalibrationOp() >>> beads = "Spherotech RCP-30-5A Lot AA01-AA04, AB01, AB02, AC01, GAA01-R" >>> bead_op.beads = flow.BeadCalibrationOp.BEADS[beads] >>> bead_op.units = {"Pacific Blue-A" : "MEBFP", ... "FITC-A" : "MEFL", ... "PE-Tx-Red-YG-A" : "MEPTR"} >>> >>> bead_op.beads_file = "tasbe/beads.fcs"
Estimate the model parameters
>>> bead_op.estimate(ex)
Plot the diagnostic plot
>>> bead_op.default_view().plot(ex)
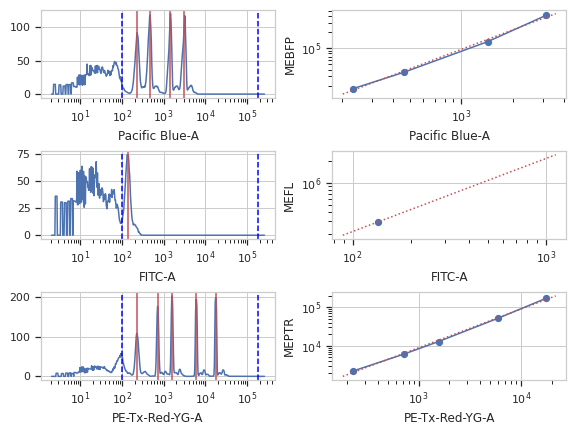
Apply the operation to the experiment
>>> ex = bead_op.apply(ex)
-
estimate(experiment)[source]¶ Estimate the calibration coefficients from the beads file.
Parameters: experiment (Experiment) – The experiment used to compute the calibration.
-
apply(experiment)[source]¶ Applies the bleedthrough correction to an experiment.
Parameters: experiment (Experiment) – the experiment to which this operation is applied Returns: A new experiment with the specified channels calibrated in physical units. The calibrated channels also have new metadata: - bead_calibration_fn : Callable (pandas.Series –> pandas.Series)
- The function to calibrate raw data to bead units
- bead_units : String
- The units this channel was calibrated to
Return type: Experiment
-
default_view(**kwargs)[source]¶ Returns a diagnostic plot to see if the peak finding is working.
Returns: An diagnostic view, call plot()to see the diagnostic plotsReturn type: IView
-
BEADS= {'Spherotech RCP-30-5A Lot AA01-AA04, AB01, AB02, AC01, GAA01-R': {'MEAP': [587, 2433, 6720, 17962, 30866, 51704, 146080], 'MEAPCY7': [718, 1920, 5133, 9324, 14210, 26735], 'MEBFP': [700, 1705, 4262, 17546, 35669, 133387, 412089], 'MECSB': [179, 400, 993, 3203, 6083, 17777, 36331], 'MECY': [1437, 4693, 12901, 36837, 76621, 261671, 1069858], 'MEFL': [692, 2192, 6028, 17493, 35674, 126907, 290983], 'MEPCY7': [32907, 107787, 503797], 'MEPE': [505, 1777, 4974, 13118, 26757, 94930, 250470], 'MEPTR': [207, 750, 2198, 6063, 12887, 51686, 170219]}, 'Spherotech RCP-30-5A Lot AG01, AF02, AD04 and AAE01': {'MEAP': [373, 1079, 3633, 9896, 28189, 79831, 151008], 'MEAPCY7': [2864, 7644, 19081, 37258], 'MEBFP': [861, 1997, 5776, 15233, 45389, 152562, 396759], 'MECSB': [216, 464, 1232, 2940, 7669, 19812, 35474], 'MECY': [1614, 4035, 12025, 31896, 95682, 353225, 1077421], 'MEFL': [792, 2079, 6588, 16471, 47497, 137049, 271647], 'MEPCY7': [14916, 42336, 153840, 494263], 'MEPE': [531, 1504, 4819, 12506, 36159, 109588, 250892], 'MEPTR': [233, 669, 2179, 5929, 18219, 63944, 188785]}, 'Spherotech RCP-30-5A Lot AK02, AK03, AK04': {'MEAP': [341, 1027, 3156, 7750, 23446, 68702, 116813], 'MEAPCY7': [173, 427, 1097, 2399, 6359, 17475, 30725], 'MEBFP': [844, 1958, 5422, 13522, 42717, 153501, 420359], 'MECSB': [205, 470, 1211, 2740, 7516, 20122, 35573], 'MECY': [1414, 3809, 10852, 27904, 85866, 324106, 1040895], 'MECY7': [12752, 39057, 142958, 448890], 'MEFL': [771, 2106, 6262, 15183, 45292, 136258, 291042], 'MEPE': [487, 1474, 4516, 11260, 34341, 107608, 260461], 'MEPTR': [205, 643, 2021, 5278, 17018, 62451, 198933]}, 'Spherotech URCP-100-2H (9 peaks)': {'MEAP': [369, 749, 3426, 10413, 50013, 177490, 500257, 1252120], 'MEAPCY7': [1363, 2656, 9791, 25120, 96513, 328967, 864905, 2268931], 'MEBFP': [1957, 5579, 16005, 53621, 168302, 459809, 1581762, 4999251], 'MECSB': [989, 2959, 8277, 25524, 71603, 173069, 491388, 1171641], 'MEFL': [3531, 11373, 34643, 107265, 324936, 835306, 2517654, 6069240], 'MEPCY': [6501, 20302, 59517, 183870, 550645, 1569470, 5109318, 17854584], 'MEPCY7': [4490, 10967, 30210, 87027, 283621, 975312, 4409101, 24259524], 'MEPE': [2785, 9525, 28421, 90313, 275589, 713181, 2209251, 5738784], 'MEPTR': [1158, 4161, 12528, 41140, 130347, 344149, 1091393, 2938710]}}¶ A dictionary containing the calibrated beads that Cytoflow currently knows about. The available bead sets, the fluorophores and the laser / filter sets with which they were characterized are below:
- Spherotech RCP-30-5A Lot AK02, AK03, AK04
- MECSB (Cascade Blue, 405 –> 450/50)
- MEBFP (BFP, 405 –> 530/40)
- MEFL (Fluroscein, 488 –> 530/40)
- MEPE (Phycoerythrin, 488 –> 575/25)
- MEPTR (PE-Texas Red, 488 –> 613/20)
- MECY (Cy5, 488 –> 680/30)
- MEPCY7 (PE-Cy7, 488 –> 750 LP)
- MEAP (APC, 633 –> 665/20)
- MEAPCY7 (APC-Cy7, 635 –> 750 LP)
- Spherotech RCP-30-5A Lot AG01, AF02, AD04 and AAE01
- MECSB (Cascade Blue, 405 –> 450/50)
- MEBFP (BFP, 405 –> 530/40)
- MEFL (Fluroscein, 488 –> 530/40)
- MEPE (Phycoerythrin, 488 –> 575/25)
- MEPTR (PE-Texas Red, 488 –> 613/20)
- MECY (Cy5, 488 –> 680/30)
- MEPCY7 (PE-Cy7, 488 –> 750 LP)
- MEAP (APC, 633 –> 665/20)
- MEAPCY7 (APC-Cy7, 635 –> 750 LP)
- Spherotech RCP-30-5A Lot AA01-AA04, AB01, AB02, AC01, GAA01-R
- MECSB (Cascade Blue, 405 –> 450/50)
- MEBFP (BFP, 405 –> 530/40)
- MEFL (Flurosceine, 488 –> 530/40)
- MEPE (Phycoerythrin, 488 –> 575/25)
- MEPTR (PE-Texas Red, 488 –> 613/20)
- MECY (Cy5, 488 –> 680/30)
- MEPCY7 (PE-Cy7, 488 –> 750 LP)
- MEAP (APC, 633 –> 665/20)
- MEAPCY7 (APC-Cy7, 635 –> 750 LP)
- Spherotech URCP-100-2H (9 peaks)
- MECSB (Cascade Blue, 405 –> 450/50)
- MEBFP (BFP, 405 –> 530/40)
- MEFL (Flurosceine, 488 –> 530/40)
- MEPE (Phycoerythrin, 488 –> 575/25)
- MEPTR (PE-Texas Red, 488 –> 613/20)
- MECY (Cy5, 488 –> 680/30)
- MEPCY7 (PE-Cy7, 488 –> 750 LP)
- MEAP (APC, 633 –> 665/20)
- MEAPCY7 (APC-Cy7, 635 –> 750 LP)
- Spherotech RCP-30-5A Lot AK02, AK03, AK04
-
-
class
cytoflow.operations.bead_calibration.BeadCalibrationDiagnostic[source]¶ Bases:
traits.has_traits.HasStrictTraitsA diagnostic view for BeadCalibrationOp.
Plots the smoothed histogram of the bead data; the peak locations; a scatter plot of the raw bead fluorescence values vs the calibrated unit values; and a line plot of the model that was computed. Make sure that the relationship is linear; if it’s not, it likely isn’t a good calibration!
-
op¶ The operation instance whose parameters we’re plotting. Set automatically if you created the instance using
BeadCalibrationOp.default_view().Type: Instance(BeadCalibrationOp)
-